1. 背景:なぜ海面高度を見るのか
これまでの回では、SOI、Niño3.4、SSTを使ってENSOや長期トレンドを調べてきた。 ここでは、海面高度偏差(SLA: Sea Level Anomaly)を使う。
SSTは海面の温度を表す。一方、SLAは海面の高さの偏差であり、海水の熱膨張、風による水の吹き寄せ、 海洋循環の変化などを反映する。つまり、SSTよりも海洋の力学的な変化を見やすい場合がある。
この課題の狙いは、08のENSOコンポジットと09のトレンド解析を、SLAデータに応用することである。
2. この回の課題
以下の3つの図を作成し、SSTで得られた結果と比較する。
- SLA長期トレンドマップ
- SLAトレンドのp値マップ
- El Niño / La Niña時のSLAコンポジット
提出するもの
- 完成したPythonコード
- 作成された図(PNG)
- SSTのENSOコンポジット・SSTトレンドとの比較コメント
3. 使用データ
このページと同じディレクトリに次のファイルを置く。
| ファイル | 内容 |
|---|---|
| sla_monthly_light_2deg_1993_2025.nc | CMEMS月平均SLAを約2度格子へ軽量化したデータ。変数は sla(time, latitude, longitude)。 |
| sstoi.indices | Niño3.4 anomalyを含むENSO指数データ。 |
SLAの単位は通常 m である。図では見やすくするため、トレンドは mm/year、コンポジットは cm に変換する。
4. 計算の流れ
- SLAデータを読み込む
- Niño3.4 anomalyを読み込み、El Niño / La Niña月を判定する
- 各格子点でSLAを時間方向に線形回帰する
- 傾きとp値を保存する
- SLAトレンドマップとp値マップを描く
- El Niño月・La Niña月だけを抽出して平均する
- El Niño、La Niña、差分のSLAコンポジット図を描く
STEP 1
5. データの読み込み
import numpy as np
import pandas as pd
import xarray as xr
import matplotlib.pyplot as plt
from scipy import stats
sla_file = "______________________________.nc"
nino_file = "______________"
start_year = ______
end_year = ______
alpha = 0.05
ds = xr.open_dataset(sla_file)
sla = ds["____"]
lat = ds["________"].values
lon = ds["_________"].values
time_sla = pd.to_datetime(ds["time"].values)
sla = sla.sel(time=slice(f"{start_year}-01-01", f"{end_year}-12-31"))
time_sla = pd.to_datetime(sla["time"].values)
print(sla)ポイント
- xarray.open_dataset でNetCDFを読む
- sla は時間×緯度×経度の3次元データである
- 09のSSTトレンドと同様に、まず解析期間を指定する
STEP 2
6. Niño3.4によるENSO判定
colnames = [
"YR", "MON",
"NINO12", "ANOM12",
"NINO3", "ANOM3",
"NINO4", "ANOM4",
"NINO34", "ANOM34"
]
nino_df = pd.read_csv(
nino_file,
sep=r"\s+",
skiprows=1,
names=colnames
)
nino_df["time"] = pd.to_datetime(
dict(year=nino_df["YR"], month=nino_df["MON"], day=15)
)
nino_df = nino_df.set_index("time")
nino34 = nino_df["________"].astype(float)
nino34_smooth = nino34.rolling(
window=______,
center=True
).mean()
el_event = nino34_smooth >= ______
la_event = nino34_smooth <= -______ポイント
- 08と同じく、ENSO判定にはNiño3.4 anomalyを使う
- 5か月移動平均により、短い月ごとの揺らぎをならす
- ここでは簡略化して、5か月継続条件は使わず、5か月移動平均が±0.5℃を超える月をイベント月とする
より厳密に行う場合は、08で使った detect_runs 関数を使って「5か月以上継続」の条件を加えてもよい。
STEP 3
7. SLAの長期トレンドとp値
09で行ったSSTトレンド解析と同じ考え方で、各格子点におけるSLAの時間変化を線形回帰する。
years = time_sla.year + (time_sla.month - ______) / 12.0
years = np.asarray(years, dtype=float)
def calc_trend_pvalue(x, y):
x = np.asarray(x, dtype=float)
y = np.asarray(y, dtype=float)
mask = np.isfinite(x) & np.isfinite(y)
x = x[mask]
y = y[mask]
n = len(x)
if n < 3:
return np.nan, np.nan, np.nan, np.nan
slope, intercept = np.polyfit(x, y, 1)
r = np.corrcoef(x, y)[0, 1]
yhat = slope * x + intercept
residual = y - yhat
dof = n - 2
sse = np.sum(residual**2)
sxx = np.sum((x - np.mean(x))**2)
mse = sse / dof
se_slope = np.sqrt(mse / sxx)
tval = slope / se_slope
pval = 2 * (1 - stats.t.cdf(np.abs(tval), df=dof))
return slope, pval, r, nポイント
- slope はSLAのトレンドで、単位は m/year
- pval は「傾きが0である」という帰無仮説に対するp値
- p値が小さいことと、物理的に重要であることは同じではない
STEP 4
8. トレンドマップとp値マップの描画
sla_np = sla.values
ntime, nlat, nlon = sla_np.shape
trend = np.full((nlat, nlon), np.nan)
pval = np.full((nlat, nlon), np.nan)
for i in range(nlat):
for j in range(nlon):
y = sla_np[:, i, j]
slope, p, r, n = calc_trend_pvalue(_____, _____)
trend[i, j] = _____
pval[i, j] = _____
sig_mask = pval < alpha
trend_mm = trend * ______
def plot_global_map(data, title, cbar_label, save_name,
cmap="RdBu_r", vmin=None, vmax=None, sig=None):
fig, ax = plt.subplots(figsize=(14, 6))
pcm = ax.pcolormesh(
lon, lat, data,
shading="auto",
cmap=cmap,
vmin=vmin,
vmax=vmax
)
ax.set_xlim(-180, 180)
ax.set_ylim(-80, 80)
ax.set_xlabel("Longitude")
ax.set_ylabel("Latitude")
ax.set_title(title)
ax.grid(True, alpha=0.3)
cbar = fig.colorbar(pcm, ax=ax, pad=0.02)
cbar.set_label(cbar_label)
if sig is not None:
yy, xx = np.where(sig)
ax.scatter(lon[xx], lat[yy], s=2, c="k", alpha=0.45, label=f"p < {alpha}")
ax.legend(loc="lower left")
plt.tight_layout()
plt.savefig(save_name, dpi=300, bbox_inches="tight", facecolor="white")
plt.show()ポイント
- SLAトレンドは m/year で出るので、図では 1000 を掛けて mm/year にする
- sig_mask はp値が0.05未満の場所を表す
- 黒点が多い場合、「p値が小さい場所が多い」と言えるが、「すべて物理的に重要」とは言えない
STEP 5
9. ENSOコンポジットの計算
enso_df = pd.DataFrame({
"nino34": nino34,
"nino34_smooth": nino34_smooth,
"el": el_event,
"la": la_event
}).dropna()
tmp = enso_df.reindex(time_sla)
el_mask = tmp["el"].fillna(False).values
la_mask = tmp["la"].fillna(False).values
sla_el = sla.isel(time=________).mean(dim="time", skipna=True)
sla_la = sla.isel(time=________).mean(dim="time", skipna=True)
sla_diff = ________ - ________
sla_el_cm = sla_el.values * ______
sla_la_cm = sla_la.values * ______
sla_diff_cm = sla_diff.values * ______ポイント
- 08と同じく、ENSOイベント月だけを抽出して平均する
- SLAは m なので、図では 100 を掛けて cm にする
- sla_diff は El Niño と La Niña の差分である
STEP 6
10. ENSOコンポジット図の描画
fig, axes = plt.subplots(3, 1, figsize=(14, 15), constrained_layout=True)
panels = [
(sla_el_cm, "El Niño composite SLA", "SLA (cm)", -15, 15),
(sla_la_cm, "La Niña composite SLA", "SLA (cm)", -15, 15),
(sla_diff_cm, "El Niño - La Niña SLA", "SLA difference (cm)", -20, 20),
]
for ax, (data, title, label, vmin, vmax) in zip(axes, panels):
pcm = ax.pcolormesh(
lon, lat, data,
shading="auto",
cmap="________",
vmin=vmin,
vmax=vmax
)
ax.set_xlim(-180, 180)
ax.set_ylim(-80, 80)
ax.set_xlabel("Longitude")
ax.set_ylabel("Latitude")
ax.set_title(title)
ax.grid(True, alpha=0.3)
cbar = fig.colorbar(pcm, ax=ax, pad=0.02)
cbar.set_label(label)
plt.savefig("sla_enso_composite.png", dpi=300, bbox_inches="tight", facecolor="white")
plt.show()ポイント
- 正負の偏差を見るため、赤青の発散カラーマップを使う
- El NiñoとLa Niñaでは、赤道太平洋のSLA分布が反転する
- 差分図を見ると、ENSOにともなう海面高度変化が最もわかりやすい
11. 考察課題:SSTとSLAは何が違うか
作成したSLAのENSOコンポジット、SLA長期トレンド、p値マップを見て、 これまでに作成したSSTのENSOコンポジットおよびSST長期トレンドと比較しなさい。
考察せよ
- El Niño − La Niña の差分図について、SSTとSLAで似ている海域・異なる海域を考察せよ。
- 赤道太平洋において、SSTとSLAがどのように変化しているかを考察せよ。
- SLAトレンドは全球で一様に上昇しているか。地域差がある場合、その特徴を考察せよ。
- SSTトレンドとSLAトレンドの空間分布は一致しているか。一致しない場合、その理由を考察せよ。
- p値が小さい格子点が多い場合、それは「物理的に重要である」ことを意味するのか考察せよ。
注意
p値は「変化量の大きさ」を表しているわけではない。 変化量の大きさはトレンドの値を見て判断し、その値が統計的に0と区別できるかどうかをp値で判断する。
p値は「変化量の大きさ」を表しているわけではない。 変化量の大きさはトレンドの値を見て判断し、その値が統計的に0と区別できるかどうかをp値で判断する。
12. 解答の表示
課題提出後、確認用としてパスワードを入力すると、完全スクリプトと作成例の図を表示する。
穴埋めの要点
sla_monthly_light_2deg_1993_2025
sstoi.indices
1993
2024
sla
latitude
longitude
ANOM34
5
0.5
0.5
0.5
years, y
slope
p
1000
el_mask
la_mask
sla_el, sla_la
100
100
100
RdBu_r作成例の図
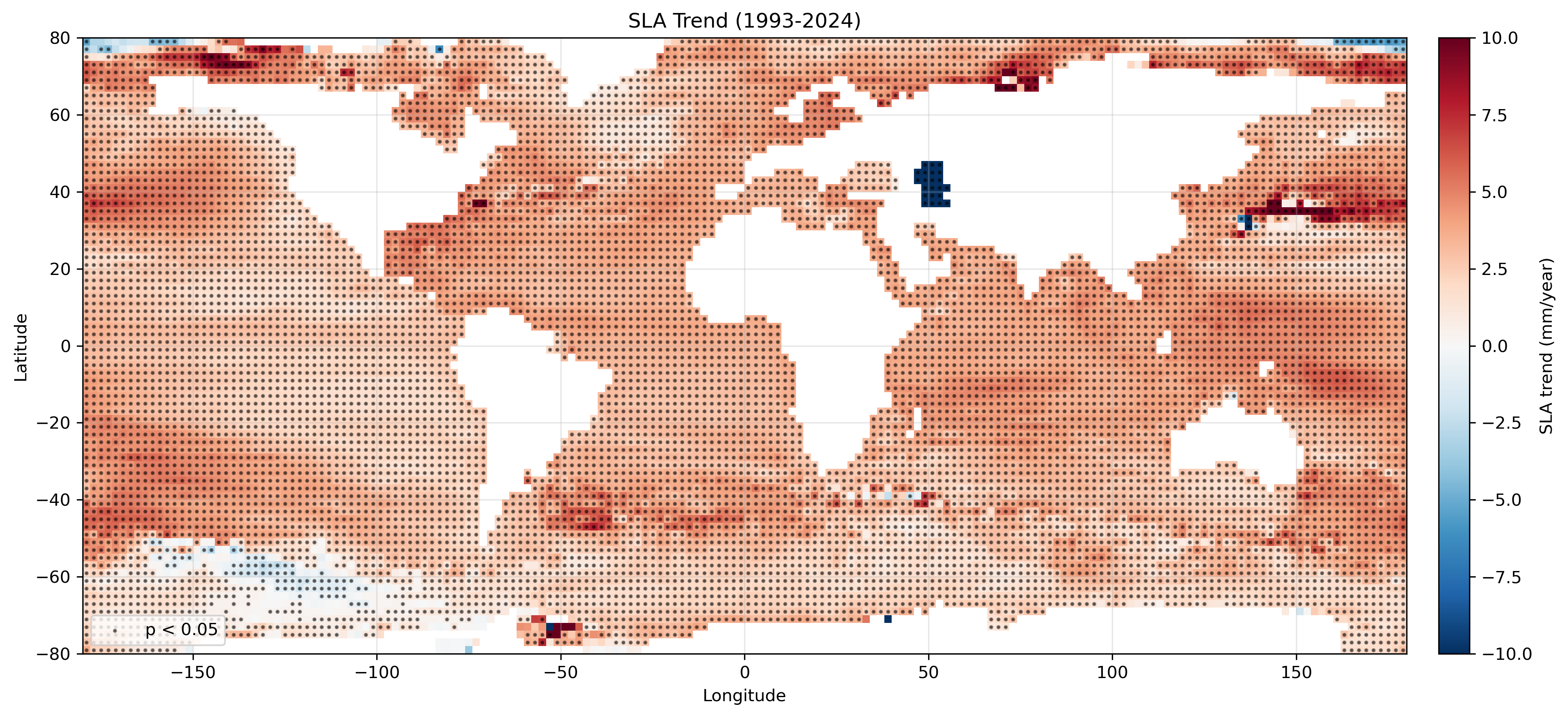
SLA長期トレンドの作成例。単位は mm/year。黒点は p < 0.05 の格子点を示す。
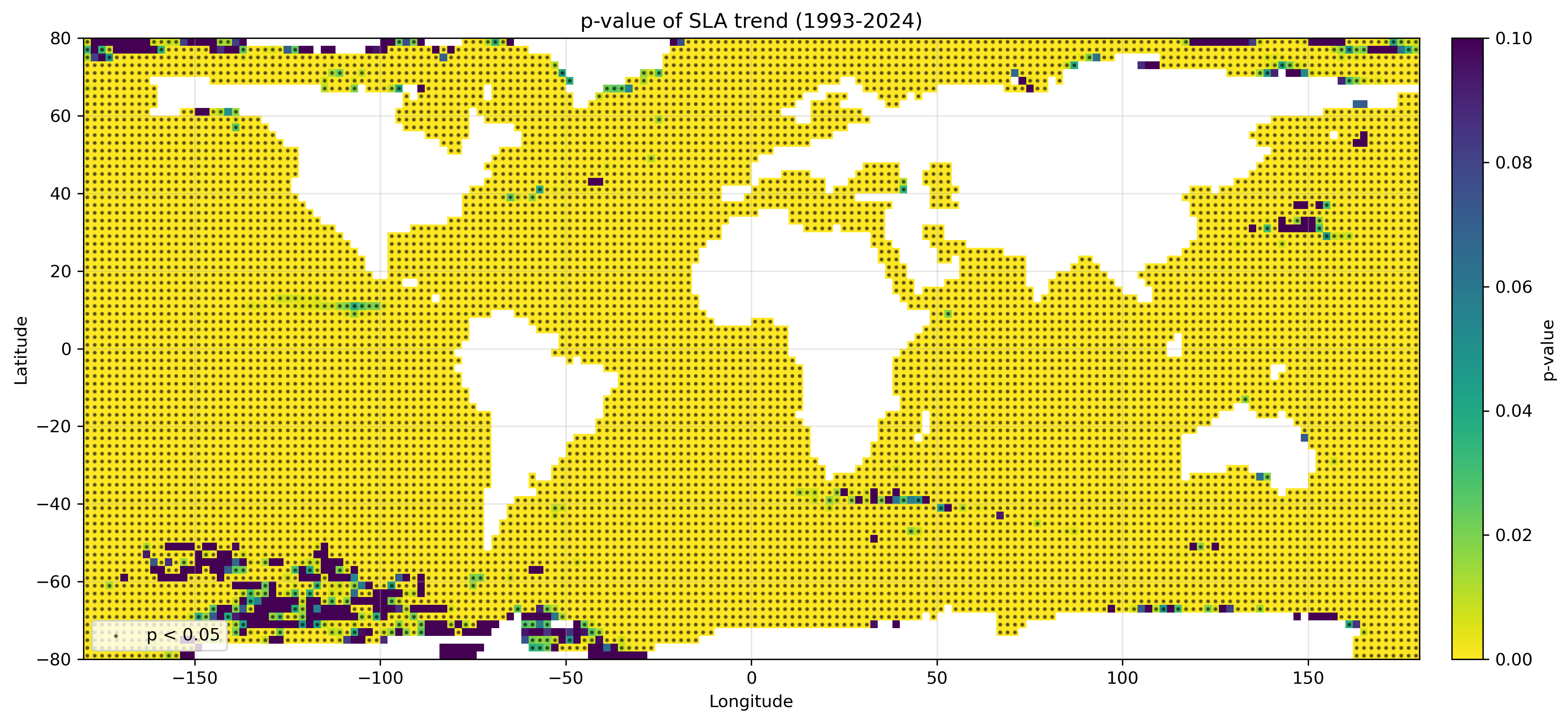
SLAトレンドに対するp値マップの作成例。
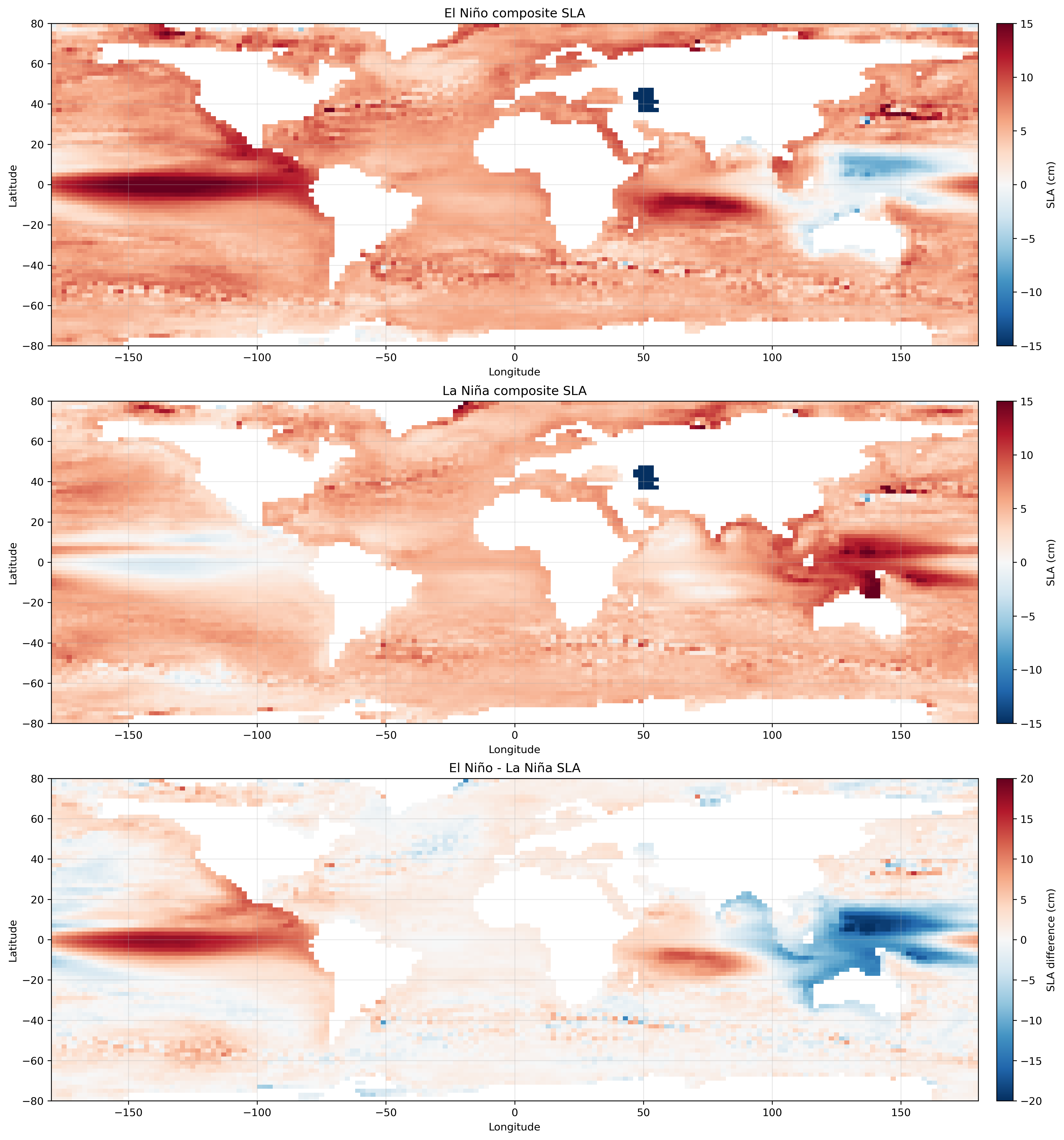
El Niño月、La Niña月、El Niño−La NiñaのSLAコンポジットの作成例。
完全スクリプト
import numpy as np
import pandas as pd
import xarray as xr
import matplotlib.pyplot as plt
from scipy import stats
sla_file = "sla_monthly_light_2deg_1993_2025.nc"
nino_file = "sstoi.indices"
start_year = 1993
end_year = 2024
alpha = 0.05
nino_threshold = 0.5
smooth_window = 5
ds = xr.open_dataset(sla_file)
sla = ds["sla"]
lat = ds["latitude"].values
lon = ds["longitude"].values
time_sla = pd.to_datetime(ds["time"].values)
sla = sla.sel(time=slice(f"{start_year}-01-01", f"{end_year}-12-31"))
time_sla = pd.to_datetime(sla["time"].values)
colnames = [
"YR", "MON",
"NINO12", "ANOM12",
"NINO3", "ANOM3",
"NINO4", "ANOM4",
"NINO34", "ANOM34"
]
nino_df = pd.read_csv(
nino_file,
sep=r"\s+",
skiprows=1,
names=colnames
)
nino_df["time"] = pd.to_datetime(
dict(year=nino_df["YR"], month=nino_df["MON"], day=15)
)
nino_df = nino_df.set_index("time")
nino34 = nino_df["ANOM34"].astype(float)
nino34_smooth = nino34.rolling(
window=smooth_window,
center=True
).mean()
el_event = nino34_smooth >= nino_threshold
la_event = nino34_smooth <= -nino_threshold
years = time_sla.year + (time_sla.month - 0.5) / 12.0
years = np.asarray(years, dtype=float)
def calc_trend_pvalue(x, y):
x = np.asarray(x, dtype=float)
y = np.asarray(y, dtype=float)
mask = np.isfinite(x) & np.isfinite(y)
x = x[mask]
y = y[mask]
n = len(x)
if n < 3:
return np.nan, np.nan, np.nan, np.nan
slope, intercept = np.polyfit(x, y, 1)
r = np.corrcoef(x, y)[0, 1]
yhat = slope * x + intercept
residual = y - yhat
dof = n - 2
sse = np.sum(residual**2)
sxx = np.sum((x - np.mean(x))**2)
mse = sse / dof
se_slope = np.sqrt(mse / sxx)
tval = slope / se_slope
pval = 2 * (1 - stats.t.cdf(np.abs(tval), df=dof))
return slope, pval, r, n
sla_np = sla.values
ntime, nlat, nlon = sla_np.shape
trend = np.full((nlat, nlon), np.nan)
pval = np.full((nlat, nlon), np.nan)
rmap = np.full((nlat, nlon), np.nan)
nmap = np.full((nlat, nlon), np.nan)
for i in range(nlat):
for j in range(nlon):
y = sla_np[:, i, j]
slope, p, r, n = calc_trend_pvalue(years, y)
trend[i, j] = slope
pval[i, j] = p
rmap[i, j] = r
nmap[i, j] = n
sig_mask = pval < alpha
trend_mm = trend * 1000.0
def plot_global_map(data, title, cbar_label, save_name,
cmap="RdBu_r", vmin=None, vmax=None, sig=None):
fig, ax = plt.subplots(figsize=(14, 6))
pcm = ax.pcolormesh(
lon, lat, data,
shading="auto",
cmap=cmap,
vmin=vmin,
vmax=vmax
)
ax.set_xlim(-180, 180)
ax.set_ylim(-80, 80)
ax.set_xlabel("Longitude")
ax.set_ylabel("Latitude")
ax.set_title(title)
ax.grid(True, alpha=0.3)
cbar = fig.colorbar(pcm, ax=ax, pad=0.02)
cbar.set_label(cbar_label)
if sig is not None:
yy, xx = np.where(sig)
ax.scatter(lon[xx], lat[yy], s=2, c="k", alpha=0.45, label=f"p < {alpha}")
ax.legend(loc="lower left")
plt.tight_layout()
plt.savefig(save_name, dpi=300, bbox_inches="tight", facecolor="white")
plt.show()
plot_global_map(
trend_mm,
title=f"SLA Trend ({start_year}-{end_year})",
cbar_label="SLA trend (mm/year)",
save_name="sla_trend_1993_2024.png",
cmap="RdBu_r",
vmin=-10,
vmax=10,
sig=sig_mask
)
plot_global_map(
pval,
title=f"p-value of SLA trend ({start_year}-{end_year})",
cbar_label="p-value",
save_name="sla_pvalue_1993_2024.png",
cmap="viridis_r",
vmin=0,
vmax=0.1,
sig=sig_mask
)
enso_df = pd.DataFrame({
"nino34": nino34,
"nino34_smooth": nino34_smooth,
"el": el_event,
"la": la_event
}).dropna()
tmp = enso_df.reindex(time_sla)
el_mask = tmp["el"].fillna(False).values
la_mask = tmp["la"].fillna(False).values
sla_el = sla.isel(time=el_mask).mean(dim="time", skipna=True)
sla_la = sla.isel(time=la_mask).mean(dim="time", skipna=True)
sla_diff = sla_el - sla_la
sla_el_cm = sla_el.values * 100.0
sla_la_cm = sla_la.values * 100.0
sla_diff_cm = sla_diff.values * 100.0
fig, axes = plt.subplots(3, 1, figsize=(14, 15), constrained_layout=True)
panels = [
(sla_el_cm, "El Niño composite SLA", "SLA (cm)", -15, 15),
(sla_la_cm, "La Niña composite SLA", "SLA (cm)", -15, 15),
(sla_diff_cm, "El Niño - La Niña SLA", "SLA difference (cm)", -20, 20),
]
for ax, (data, title, label, vmin, vmax) in zip(axes, panels):
pcm = ax.pcolormesh(
lon, lat, data,
shading="auto",
cmap="RdBu_r",
vmin=vmin,
vmax=vmax
)
ax.set_xlim(-180, 180)
ax.set_ylim(-80, 80)
ax.set_xlabel("Longitude")
ax.set_ylabel("Latitude")
ax.set_title(title)
ax.grid(True, alpha=0.3)
cbar = fig.colorbar(pcm, ax=ax, pad=0.02)
cbar.set_label(label)
plt.savefig("sla_enso_composite.png", dpi=300, bbox_inches="tight", facecolor="white")
plt.show()